Input data and parameters
Input
| Analysis date: | Fri Jul 18 01:50:08 GMT 2025 |
| BAM file: | DE-4.markdup.sorted.bam |
| Counting algorithm: | uniquely-mapped-reads |
| GTF file: | genes.filtered.gtf |
| Number of bases for 5'-3' bias computation: | 100 |
| Number of transcripts for 5'-3' bias computation: | 1,000 |
| Paired-end sequencing: | yes |
| Protocol: | strand-specific-reverse |
| Sorting performed: | yes |
Summary
Reads alignment
| Number of mapped reads (left/right): | 93,948,658 / 93,926,934 |
| Number of aligned pairs (without duplicates): | 93,895,752 |
| Total number of alignments: | 208,586,248 |
| Number of secondary alignments: | 20,710,656 |
| Number of non-unique alignments: | 30,022,681 |
| Aligned to genes: | 90,568,015 |
| Ambiguous alignments: | 656,686 |
| No feature assigned: | 87,330,852 |
| Missing chromosome in annotation: | 8,014 |
| Not aligned: | 0 |
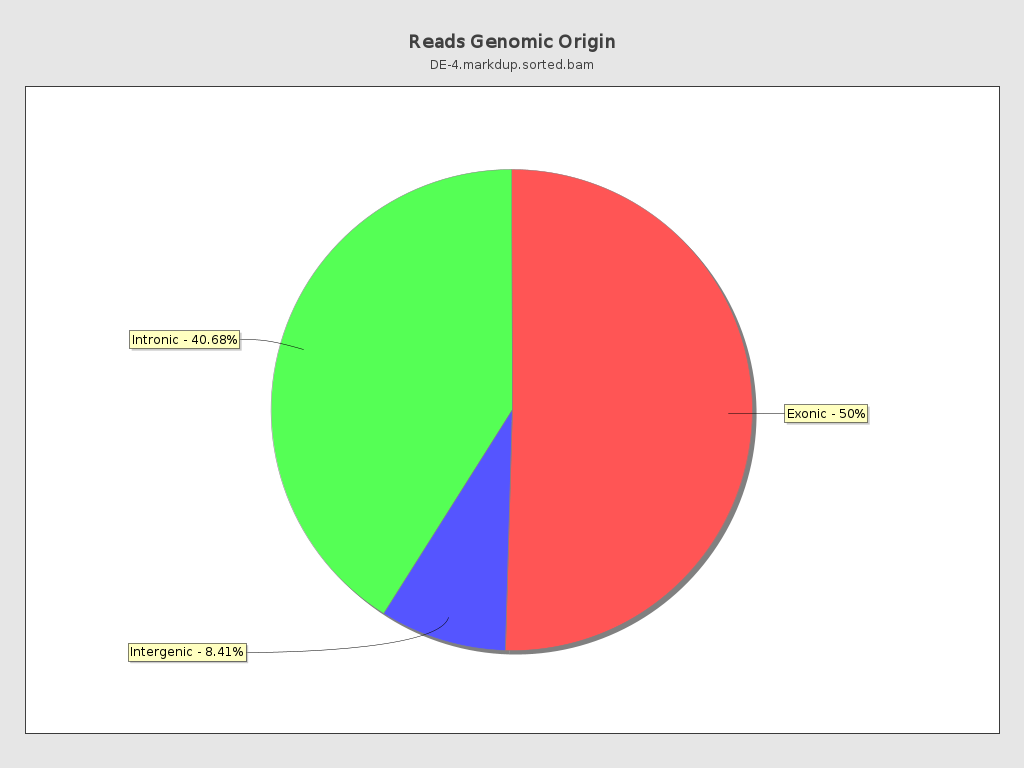
Reads genomic origin
| Exonic: | 90,568,015 / 50.91% |
| Intronic: | 72,364,513 / 40.68% |
| Intergenic: | 14,966,339 / 8.41% |
| Intronic/intergenic overlapping exon: | 4,799,963 / 2.7% |
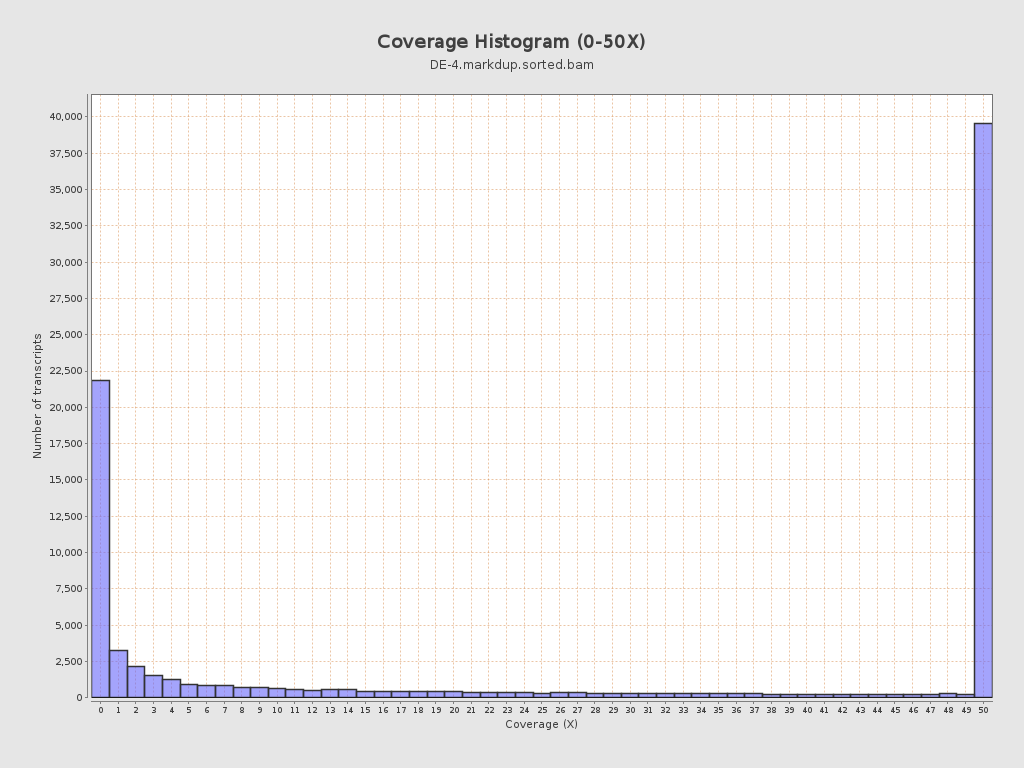
Transcript coverage profile
| 5' bias: | 0.59 |
| 3' bias: | 0.37 |
| 5'-3' bias: | 1.37 |
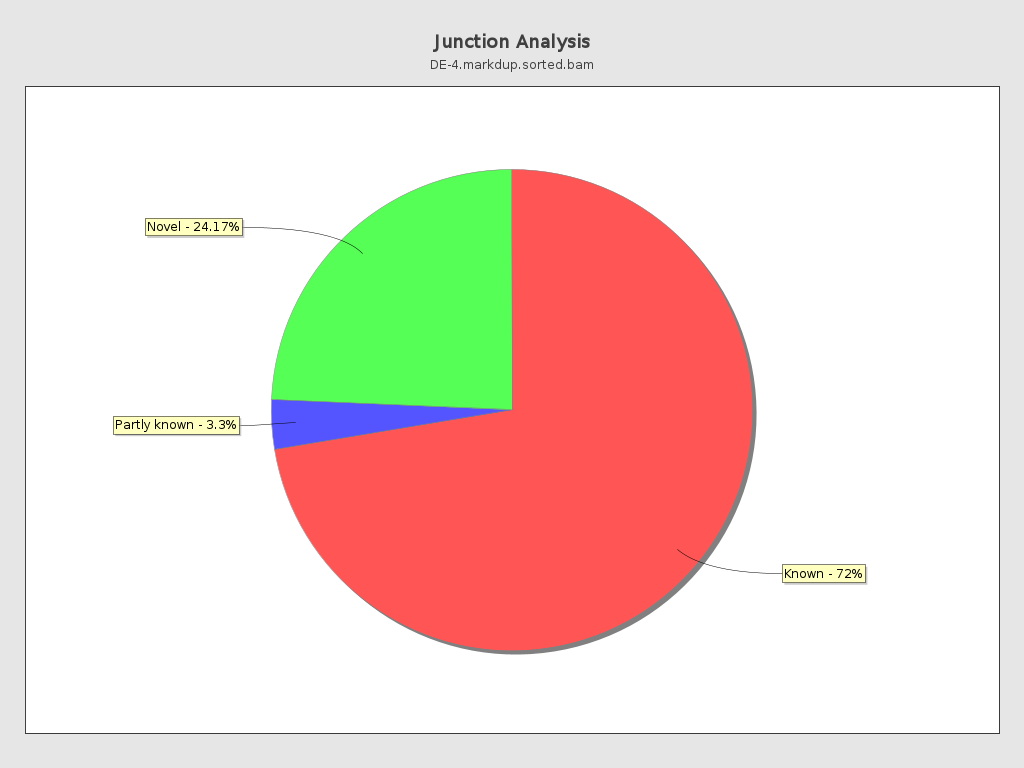
Junction analysis
| Reads at junctions: | 52,511,629 |
| AGGT | 8.18% |
| ACCT | 5.22% |
| TCCT | 3.37% |
| AGGA | 3.36% |
| ATCT | 3.17% |
| AGCT | 2.98% |
| AGGC | 2.67% |
| GCCT | 2.62% |
| CCCT | 2.34% |
| AGAT | 2.16% |
| AGGG | 2.1% |

.png)
.png)
.png)